Your haplogroup H1a emerged approximately 12,200 years ago
Your root haplogroup H emerged approximately 12,000 years ago in Western Eurasia, particularly in Europe and the Near East
Confidence Level: Medium-Low
This prediction has been made with medium to low confidence due to the quality of available genetic markers. Results should be interpreted with caution.
About Haplogroup H
Detailed information about your maternal lineage and genetic heritage
Haplogroup Overview
Haplogroup H1 is one of the most prominent subclades of mitochondrial DNA (mtDNA) haplogroup H, which itself is the most common maternal lineage in Europe. Haplogroup H1 is particularly significant in the genetic history of European populations, as well as some populations in North Africa and the Near East. H1 is maternally inherited and plays an essential role in understanding the migration, adaptation, and genetic diversity of early human populations in Europe following the Last Glacial Maximum (LGM).
Origin and Evolution
Haplogroup H1 is estimated to have originated approximately 13,000 to 15,000 years ago, most likely in the Iberian Peninsula or southern France. It is a descendant of haplogroup H, which emerged during or after the Last Glacial Maximum, around 20,000 to 25,000 years ago. The origin of H1 is closely tied to the post-glacial recolonization of Europe when populations that had retreated to glacial refugia in southern Europe began migrating northward as the climate warmed and the ice sheets retreated.
During the LGM, large portions of Europe were uninhabitable due to the extensive ice cover. Populations in Europe were forced to take refuge in warmer areas, particularly in southern regions such as Iberia, Italy, and the Balkans. Haplogroup H1 likely expanded from these refugia as humans migrated back into Central and Northern Europe during the Holocene, around 10,000 years ago.
Subclades and Geographic Distribution
Haplogroup H1 is one of the largest and most widespread subclades of haplogroup H, with numerous branches and subclades found in different parts of Europe, North Africa, and the Near East. Some of the significant subclades of H1 include:
H1a: This subclade is distributed across Europe, particularly in Western and Central Europe, but is also found in North Africa and the Near East. Its distribution reflects the migrations and expansions of populations during the post-glacial period.
H1b: Found primarily in Eastern and Central Europe, H1b is also present in Scandinavia and some parts of Western Europe. This subclade may be associated with the expansion of early farming communities during the Neolithic period.
H1c: Found mainly in Central and Eastern Europe, H1c provides evidence for the migration patterns of populations in these regions during prehistoric times.
H1f: This subclade is found at low frequencies in Europe and may reflect more localized population movements within the continent.
Geographic Distribution and Significance
Europe
Haplogroup H1 is widespread across Europe, with the highest frequencies found in the Iberian Peninsula, particularly in Spain and Portugal. It is also prevalent in France, Italy, and parts of the British Isles, including Ireland. The presence of H1 in these regions reflects its origin in Iberia and southern France, where populations carrying H1 likely expanded after the Last Glacial Maximum.
H1 is also found in significant frequencies in Central and Eastern Europe, where it contributed to the recolonization of these regions as populations moved northward from southern Europe. Its presence in Scandinavia, including Norway, Sweden, and Denmark, reflects the later migrations of populations into northern Europe.
In addition, H1 is particularly common in the Basque population of northern Spain and southwestern France, where it is found at some of the highest frequencies in Europe. The Basques are thought to have preserved a relatively ancient genetic makeup due to their relative isolation from other European populations, and the high frequency of haplogroup H1 among the Basques suggests that they are one of the modern groups most closely related to the original populations that expanded from Iberia after the Ice Age.
North Africa
Haplogroup H1 is also found in relatively high frequencies in North Africa, particularly among Berber populations in Morocco, Algeria, and Tunisia. This distribution likely reflects ancient migrations from Europe to North Africa during the post-glacial period and subsequent gene flow between these regions.
The shared genetic heritage between European and North African populations suggests that human populations have been moving and interacting across the Mediterranean for thousands of years. The high frequency of H1 among Berbers highlights the historical genetic connections between southern Europe and North Africa.
Near East
In the Near East, haplogroup H1 is present at lower frequencies compared to Europe, but its presence is significant in populations such as the Druze in Lebanon and Israel. The distribution of H1 in the Near East likely reflects ancient population movements and trade routes connecting Europe with the Near East during prehistoric times and later historical periods.
Central Asia
Though much less common, haplogroup H1 has also been detected in some populations of Central Asia. This likely reflects ancient migration routes from Europe into Asia, including during the Bronze Age and later periods of nomadic expansion.
Population Genetics and Historical Insights
Haplogroup H1 has been extensively studied in the context of European population genetics, particularly regarding the post-glacial recolonization of Europe. Its distribution provides critical evidence for the migration routes taken by early humans as they moved out of glacial refugia and resettled the continent.
Haplogroup H1 is also a key marker in studies of ancient DNA, where it has been found in prehistoric human remains across Europe, including Mesolithic hunter-gatherers and Neolithic farming communities. These findings show that H1 was present in Europe before the arrival of agriculture and continued to be a dominant maternal lineage during the transition to farming.
The widespread presence of haplogroup H1 across Europe and North Africa reflects the long history of human migration and interaction between these regions. Genetic studies have used H1 to trace the movements of populations across the Mediterranean and into Europe during different periods, including the Ice Age, the Neolithic expansion of farming, and later historical migrations.
Ancient DNA Studies
Haplogroup H1 has been identified in several ancient DNA studies, providing direct evidence of its presence in prehistoric populations. For example, H1 has been found in Mesolithic hunter-gatherer remains from Iberia and France, showing that it was present in Europe before the arrival of agriculture. It has also been detected in Neolithic remains from farming communities, highlighting its role in the genetic makeup of early European farmers.
Ancient DNA studies have also shown that haplogroup H1 was present in the Bell Beaker culture, which spread across Western and Central Europe during the Bronze Age. The presence of H1 in ancient DNA samples supports the theory that populations carrying this lineage played a significant role in shaping the genetic landscape of Europe during the post-glacial and Neolithic periods.
Genetic Diversity and Evolutionary Significance
Haplogroup H1 is one of the most genetically diverse subclades of haplogroup H, with numerous subclades and branches that reflect the complex migration history of Europe. This diversity suggests that H1 underwent significant expansion during the post-glacial period, as populations spread across the continent and adapted to different environments.
The evolutionary significance of haplogroup H1 lies in its role in the recolonization of Europe after the Last Glacial Maximum. As populations carrying H1 expanded from southern refugia into northern and central Europe, they likely encountered and interbred with other groups, contributing to the genetic diversity of modern European populations.
Conclusion
Haplogroup H1 is one of the most important and widespread mtDNA subclades in Europe, with a deep history that traces back to the post-glacial recolonization of the continent. Originating around 13,000 to 15,000 years ago, likely in the Iberian Peninsula, H1 expanded across Europe as populations migrated northward following the retreat of the glaciers.
Today, haplogroup H1 is found at high frequencies in Western Europe, particularly in the Basque population, as well as in Central and Eastern Europe, North Africa, and parts of the Near East. Its presence in these regions reflects the long history of human migration and interaction across Europe and the Mediterranean.
Haplogroup H1 continues to be a key focus in population genetics and ancient DNA studies, helping to unravel the complex patterns of human migration and adaptation that have shaped modern populations across Europe, North Africa, and beyond.
Historical Context
Your haplogroup is found in various populations around the world, reflecting ancient migration patterns and population movements.
Where This Lineage Is Found
- European populations (especially in Western Europe, including the Iberian Peninsula, France, and the British Isles)
- North African populations (especially in the Berber populations)
- Some populations in the Middle East
- Some populations in Central Europe and Scandinavia
- Jewish populations, particularly Ashkenazi Jews
When in Time
Your haplogroup in the context of human history
Haplogroup H
Your mtDNA haplogroup emerged in Western Europe, with notable frequencies in the Iberian Peninsula and surrounding regions, including North Africa
Neolithic Revolution
Agriculture begins, settled communities form
Bronze Age
Metalworking, writing, and early civilizations
Iron Age
Iron tools, expanded trade networks
Classical Antiquity
Greek and Roman civilizations flourish
Present Day
Modern era
Your Maternal Journey
Trace your maternal lineage through thousands of years of human history
Migrations of Your Maternal Line
About Your Maternal Journey
If every person living today could trace their maternal line back over thousands of generations, all of our lines would meet at a single woman who lived in eastern Africa between 150,000 and 200,000 years ago. Though she was one of perhaps thousands of women alive at the time, only the diverse branches of her haplogroup have survived to today. The story of your maternal line begins with her.
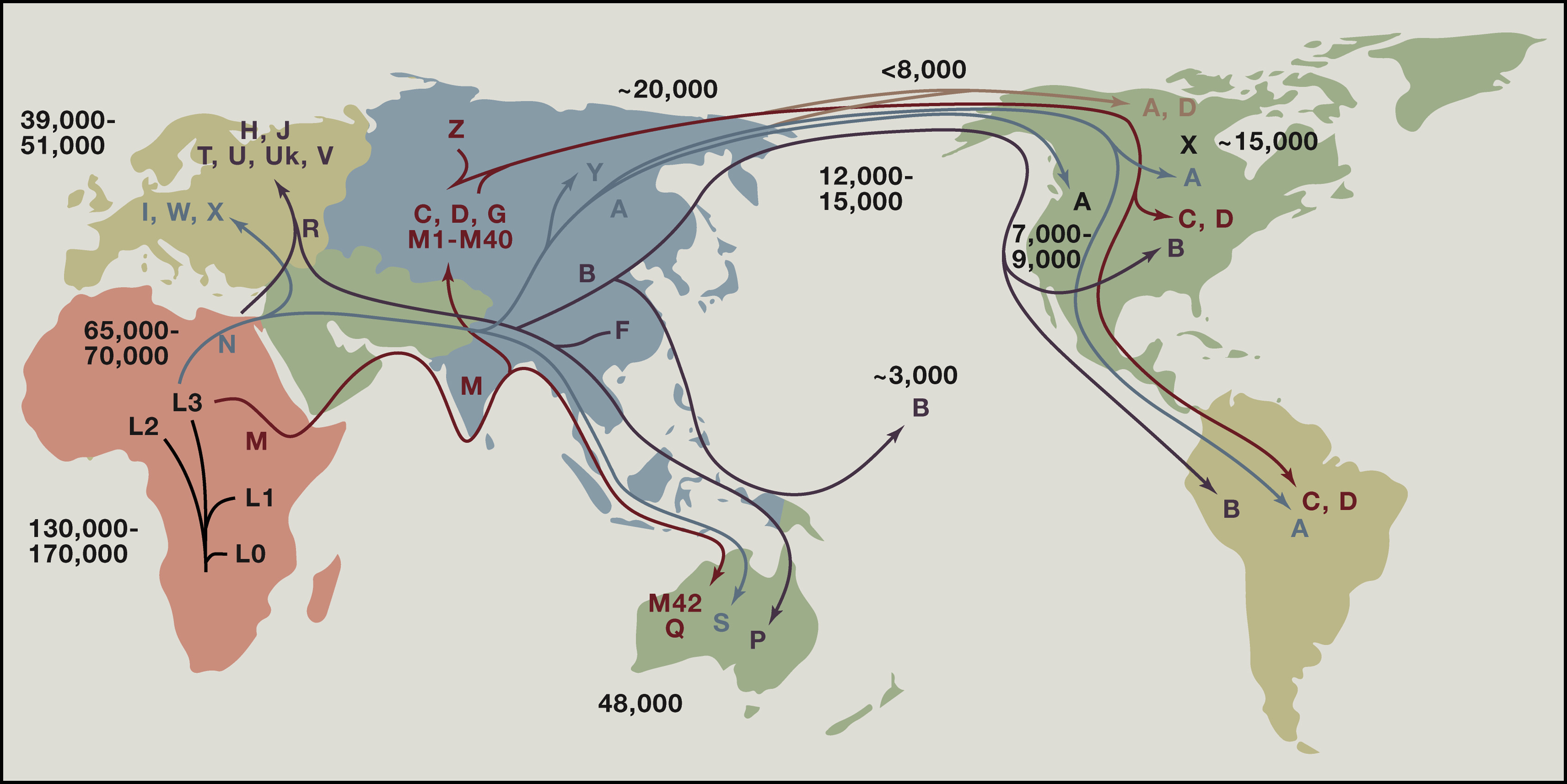
Origin and Migrations of Haplogroup H
Your maternal line stems from a branch of haplogroup H that emerged approximately 12,000 years ago in Western Europe, with notable frequencies in the Iberian Peninsula and surrounding regions, including North Africa.
Population Distribution
Where your haplogroup is found today and its connections to ancient migrations
Geographic Presence
Your maternal haplogroup H has spread across various regions through thousands of years of human migration. Its distribution today reflects both ancient population movements and more recent demographic changes.
- European populations (especially in Western Europe, including the Iberian Peninsula, France, and the British Isles)
- North African populations (especially in the Berber populations)
- Some populations in the Middle East
Frequency Statistics
Based on our database of genetic samples, your haplogroup appears with the following frequency:
- 1 in 213 of our users share your haplogroup assignment
- Your haplogroup emerged approximately 12,200 years ago
- This represents a continuous maternal lineage spanning countless generations
Modern Connections
People who carry your maternal haplogroup today are connected through an unbroken chain of mothers and daughters stretching back thousands of years. This shared lineage transcends modern national boundaries and cultural divisions.
Cultures Linked to Your Haplogroup
Ancient cultures associated with your maternal lineage based on archaeological DNA evidence
Cultural Heritage
These ancient cultures have been linked to your haplogroup H based on matching ancient DNA samples from archaeological excavations.
Primary Cultures Exact Matches
Ancient individuals with your exact haplogroup assignment belonged to these cultures.
Ancient Samples Matching Your Haplogroup
Discover ancient individuals from archaeological sites who share your maternal lineage
Ancient DNA Matches
These ancient individuals from archaeological excavations share your mtDNA haplogroup or closely related subclades. Matches are ranked by haplogroup similarity: exact matches first, followed by same base haplogroup, then close subclades.
| Portrait | Sample | Country | Era | Date | Culture | mtDNA | Match |
|---|---|---|---|---|---|---|---|

|
I1690
|
Israel | Natufian Culture in Israel - Israel - Raqefet Cave | 12000 BCE - 9500 BCE | Levantine | H |
Exact |

|
I3883
|
Iraq | Shanidar Cave, Iraq - Iraq - Shanidar | 8456 BCE - 8251 BCE | Neanderthal | H |
Exact |

|
I11270
|
Bulgaria | Neolithic Bulgaria - Bulgaria - Dzhulyunitsa (Veliko Tarnovo) | 6100 BCE - 5450 BCE | Balkan Neolithic | H |
Exact |

|
I13162
|
Serbia | Early Neolithic Starčevo Culture, Serbia - Serbia - Donja Strana-Velesnica (Bor District, Kladovo Municipality, Velesnica) | 6081 BCE - 5912 BCE | Neolithic European | H |
Exact |

|
I2373
|
Hungary | Early Neolithic Körös Culture, Hungary - Hungary - Törökszentmiklos Tiszapüspöki Karanycs haromag 3. lh. | 6000 BCE - 5500 BCE | European Neolithic | H |
Exact |

|
I0698
|
Bulgaria | Neolithic Bulgaria - Bulgaria - Yabalkovo. Dimitrovgrad. Haskovo | 6000 BCE - 5900 BCE | Balkan Neolithic | H |
Exact |

|
I6181
|
Romania | Starčevo Culture - Romania - Teleor-3 (Teleorman County, Măgura) | 6000 BCE - 5300 BCE | Early European Farmers | H |
Exact |

|
I2373
|
Hungary | Hungary EN Koros contam all - Törökszentmiklos Tiszapüspöki Karanycs haromag 3. lh. | 6000 BCE - 5500 BCE | H |
Exact | |

|
I3879
|
Bulgaria | Neolithic Malak Preslavets, Bulgaria - Bulgaria - Malak Preslavets | 5800 BCE - 5400 BCE | Balkan Neolithic | H |
Exact |

|
R10
|
Italy | Neolithic Italy - Italy - Grotta Continenza | 5721 BCE - 5634 BCE | Mediterranean Neolithic | H |
Exact |
Understanding Match Types
Understanding Your Mutations
Learn how to read and interpret your mtDNA mutation report
mtDNA Regions
The mtDNA mutations report shows mutations you have on three key regions of your mitochondrial DNA:
Reference Sequences
There are two different reference sequences used for interpreting mtDNA mutations:
RSRS (Reconstructed Sapiens Reference Sequence)
The RSRS is based on the deepest common maternal ancestor of all people alive today as well as several ancient humanoids.
How to read RSRS mutations
RSRS values are reported using a system that lists the ancestral value, the position, then your mutation.
rCRS (Revised Cambridge Reference Sequence)
The rCRS was established by researchers to provide a consistent reference for interpreting mtDNA sequences and is widely used in mitochondrial DNA research.
How to read rCRS mutations
Your rCRS values are reported by listing the location followed by your derived value.
How do the two reference sequences differ?
rCRS: Based on the mtDNA sequence of a single individual. It has limitations due to its reliance on a single individual's sequence and its European bias.
RSRS: Derived from the mtDNA sequences of multiple modern humans representing diverse populations. The RSRS provides a more robust reference point for mtDNA analysis, particularly in studies involving populations with genetic backgrounds outside of Europe.
Your Defining Mutations
These are the mutations that define your haplogroup H:
These mutations distinguish your maternal lineage from other branches of the human maternal tree.
Your mtDNA Mutations
Your mtDNA mutations are positions where your sequence differs from the rCRS (Revised Cambridge Reference Sequence). These mutations help identify your specific maternal lineage and can be used for genealogical comparisons.
Mutation Summary
HVR1: 1 mutation, HVR2: 0 mutations, Coding Region: 1 mutation
Your Additional Mutations
These mutations are present in your mtDNA but are not part of the defining mutations for haplogroup H. These are personal mutations that occurred in your maternal line after your haplogroup was established.
Missing Haplogroup-Defining Mutations
These mutations are expected for haplogroup H but were not detected in your mtDNA. This can occur due to back mutations, data quality variations, or incomplete coverage.
HVR1 Mutations (Positions 16001-16569):
Coding Region Mutations (Positions 575-16000):
Using Mutations in Genealogy: When comparing your DNA results to those of others in Group Projects, you can see what similarities and differences you might have with other members. This will help you pinpoint where your mutations may have occurred, and how other members fit within your family tree.
AI-Powered Analysis
Get personalized insights about your maternal lineage
AI ASSISTANT by DNAGENICS
This AI analysis is limited to your mtDNA haplogroup information only. No additional data or personal information is included in this analysis.
Try these questions:
The Science Behind Your Results
Understanding the methodology and answering common questions
Technical Details & FAQ
Understanding the science behind your mtDNA haplogroup prediction
mtDNA haplogroup prediction involves several sophisticated steps:
- Variant Analysis: Analysis of mitochondrial DNA variants in your genetic data
- Haplogroup Comparison: Comparison with established haplogroup-defining mutations
- Phylogenetic Analysis: Phylogenetic tree analysis to determine your specific branch
- Statistical Validation: Statistical validation of the prediction confidence
- Quality Assessment: Evaluation of data quality and marker coverage
Prediction confidence is influenced by several key factors:
- Marker Coverage: Number of available mtDNA markers in your data
- Data Quality: Quality of detected variants and sequencing accuracy
- Haplogroup Coverage: Coverage of haplogroup-defining mutations
- Ambiguous Results: Presence of conflicting or ambiguous markers
- Reference Data: Quality and completeness of reference databases
Your Confidence Level: Medium-Low confidence typically indicates partial marker coverage or some ambiguous results.
Migration paths are reconstructed using multiple scientific approaches:
- Archaeological Evidence: Archaeological evidence of population movements
- Geographic Distribution: Geographic distribution of haplogroup frequencies
- Ancient DNA: Ancient DNA samples from different time periods
- Climate Data: Historical climate and geographical data
- Molecular Clock: Molecular clock estimates for timing
The timeline is calibrated using molecular clock estimates and verified with ancient DNA samples from archaeological sites.
Our mtDNA analysis relies on extensive reference databases:
- PhyloTree: PhyloTree mtDNA phylogenetic tree (Build 17)
- Modern Sequences: Over 50,000 modern mtDNA sequences
- Ancient DNA: Ancient DNA samples spanning multiple time periods
- Population Studies: Population frequency data from scientific literature
- User Database: Our own user database for frequency calculations
The database is regularly updated with new research findings and published sequences to ensure accuracy.
Population frequencies are calculated using statistical methods:
Frequency = (Number of haplogroup occurrences / Total population sample size) × 100
Our calculations include:
- User Database: Current user database statistics
- Regional Studies: Regional population studies
- Academic Research: Published academic research data
- Quality Control: Rigorous quality control measures
Your Result: 1 in 213 users share your haplogroup assignment.
Testing providers have discontinued MTDNA/YDNA data provision:
As of January 2026, AncestryDNA is no longer providing MTDNA and YDNA haplogroup data in their raw DNA files for tests taken since this date. This means:
- AncestryDNA tests before January 2026: May still contain mtDNA/YDNA data if downloaded before the change
- AncestryDNA tests since January 2026: Will not contain mitochondrial or Y-chromosome DNA markers
- MyHeritage: Never provided mtDNA data (only YDNA, which is no longer available since January 2026)
- Alternative options: Consider using data from 23andMe, LivingDNA, ADNtro, TellMeGen, or ordering a dedicated mtDNA test from FamilyTreeDNA
If you tested with AncestryDNA recently and need mtDNA analysis, we recommend uploading data from another provider or ordering a dedicated mtDNA test kit.
Additional Resources
Research papers and publications related to your haplogroup







Understanding Your Results
Key takeaways from your maternal haplogroup analysis
Key Takeaways
Your haplogroup H connects you to a specific branch of the human maternal family tree that emerged approximately 12,200 years ago. This represents a direct genetic link to your maternal ancestors and their journey through human history.
What This Means For You
- Maternal Lineage: You carry genetic markers from your direct maternal line stretching back thousands of years
- Population History: Your haplogroup reflects the migration patterns of your maternal ancestors
- Genetic Diversity: Your specific haplogroup represents a unique branch of human genetic diversity
- Scientific Significance: You're part of ongoing research that continues to reshape our understanding of human evolution
- Individual Connection: This haplogroup assignment connects you directly to your maternal genetic heritage
